HiGHmed | Who we are.
HiGHmed started as a large scale medical informatics project within the Medical Informatics Initiative (MII) developing novel, interoperable solutions to increase the accessibility of medical patient data for clinical research and education. To improve patient care and to enable clinicians to make data-based and patient-centric decisions continues to be the core objective of the HiGHmed Consortium. We are building safe data integration centres and developing the technology for an efficient use of medical patient data across various sites to further research and health care.
The project pools the expertise of twelve university hospitals and medical faculties as well as other academic and industrial partners. The Luxembourg Institute of Health (LIH) is the first international partner to expand this strong network in Germany. The joint know-how will be applied in prototypical clinical use cases.
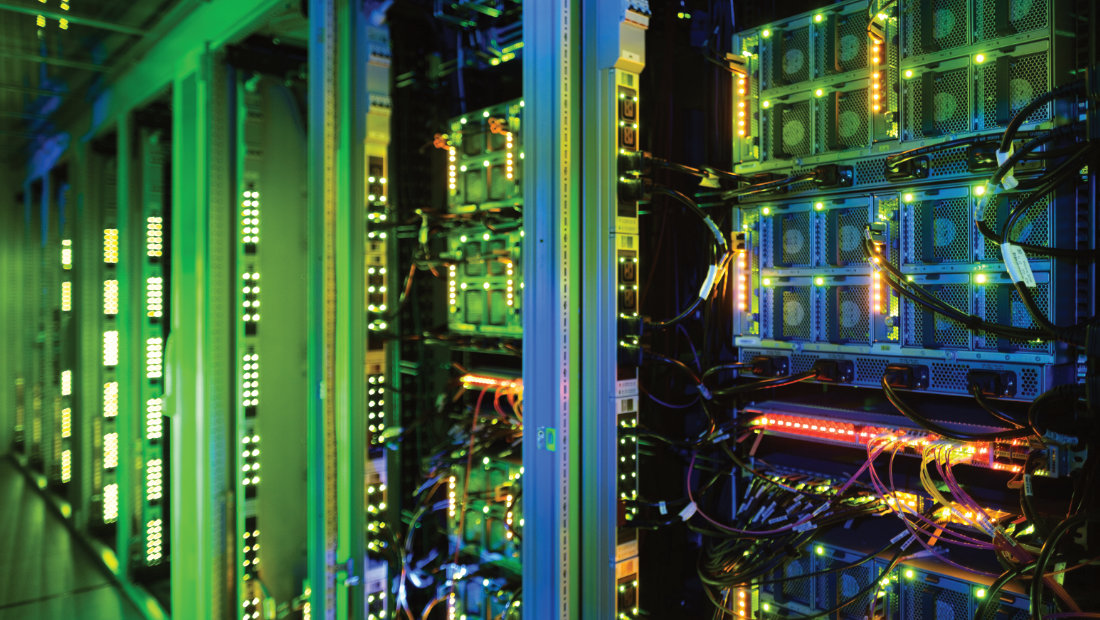
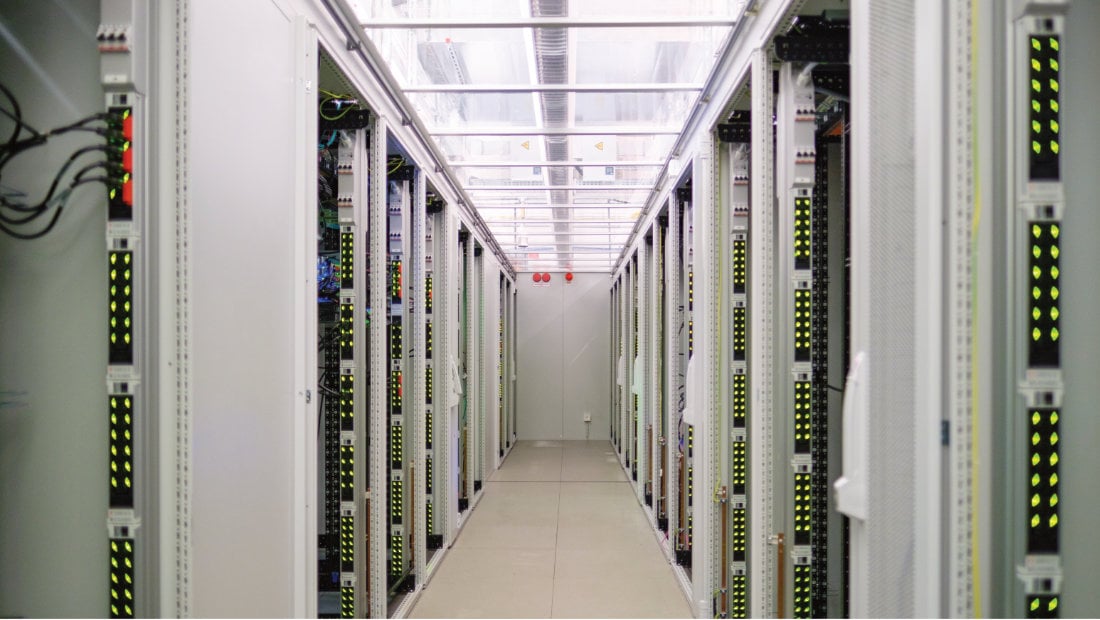



"The great challenge of building a sustainable, data-driven and patient-centered healthcare system is best met together - with progressive concepts and bold inventiveness. HiGHmed brings together numerous outstanding academic and industrial partners. Together, we are working on innovative solutions to link healthcare and research even better in the future - for the benefit of all patients."
Prof. Dr. Roland Eils
HiGHmed leader
Our partners with a Medical Data Integration Centre (MeDIC)

The Heidelberg University Hospital offers inpatients and outpatients an innovative and effective diagnosis and therapy for all complex diseases. Modern buildings with state-of-the-art equipment enable patients from all over Germany and many other countries to receive medical care that meets the highest international standards. The proximity and interlinking of the specialist departments benefit the patient: interdisciplinary cooperation ensures optimal treatment.
Partner Representative: Prof. Dr. Christoph Dieterich
Head of MeDIC: Dr. Angela Merzweiler
Deputy Head of MeDIC: Reto Wettstein
Partner within the HiGHmed Consortium:
since 2016 (founding location)
Member of the HiGHmed Association:
since 2020

Charité – Universitätsmedizin Berlin is one of the largest university hospitals in Europe, boasting 3,099 beds and approximately 100 departments and institutes spread across 4 separate campuses. At Charité, the areas of research, teaching and medical care are closely interlinked. With a total of 20,921 members of staff employed across its group of companies (17,615 of which at Charité), the organization is one of the largest employers in Berlin. Last year, Charité treated almost 125,000 in- and day case patients, in addition to over 680,000 outpatients. Charité’s Medical Faculty is one of the largest in Germany, educating and training more than 9,000 students across the subjects of medicine, dentistry, health sciences and nursing.
Partner Representative: Prof. Dr. Fabian Prasser
Deputy of Partner Representative: Martin Peuker
Head of MeDIC: Dr. Peter Brunecker
Deputy Head of MeDIC: Michael Mallach
Partner within the HiGHmed Consortium:
since 2018
Member of the HiGHmed Association:
since 2020

The University Medical Center Göttingen (UMG) - that is the Faculty of Medicine with research and teaching and the University Hospital with patient care. The bundling of these three areas under one roof is unique in southern Lower Saxony and enables numerous positive interactions, such as the prompt transfer of innovative therapy options to patient care and education.
Partner Representative: Prof. Dr. Dagmar Krefting
Head of MeDIC: Prof. Dr. Tibor Kesztyüs
Deputy Head of MeDIC: Dr. Sabine Rey
Partner within the HiGHmed Consortium:
since 2016 (founding location)
Member of the HiGHmed Association:
since 2020

As one of the largest European centers of university medicine, the University Medical Center Schleswig-Holstein (UKSH) combines top international research with interdisciplinary patient care and training of doctors of future generations. UKSH has specialists in the entire spectrum of modern medicine.
Partner Representative: Prof. Dr. Björn Bergh
Head of MeDIC: Prof. Dr. Björn Schreiweis
Deputy Head of MeDIC: Dr. Andreas Wolf
Partner within the HiGHmed Consortium:
since 2018
Member of the HiGHmed Association:
since 2020

As one of the largest European centers of university medicine, the University Medical Center Schleswig-Holstein (UKSH) combines top international research with interdisciplinary patient care and training of doctors of future generations. UKSH has specialists in the entire spectrum of modern medicine.
Partner Representative: Prof. Dr. Björn Bergh
Head of MeDIC: Prof. Dr. Björn Schreiweis
Deputy Head of MeDIC: Ann-Kristin Kock-Schoppenhauer
Partner within the HiGHmed Consortium:
since 2018
Member of the HiGHmed Association:
since 2020

The Cologne University Hospital offers medical care that is always at the cutting edge of knowledge and expertise. As a modern maximum-care hospital with around 1,500 beds, Cologne University Hospital is committed to science-oriented, innovative medicine and assumes important social tasks in research, teaching and patient care.
Partner Representative: Prof. Dr. Andreas Beyer
Deputy of Partner Representative: Prof. Dr. Oya Beyan
Head of MeDIC: René Thielsch-Nebel
Deputy Head of MeDIC: Dr. Simon Schumacher
Partner within the HiGHmed Consortium:
since 2018
Member of the HiGHmed Association:
since 2020

In cooperation with the University Hospital, the University of Münster operates a Medical Data Integration Center (MeDIC), located at the Institute of Medical Informatics. The MeDIC merges data from health research and care and offers colleagues an optimal research environment with partners of the Medical Informatics Initiative.
The analysis of data from health care leads to new insights and improves medical care. The MeDIC observes local and national data security and data protection regulations. In 2020, a quality management system in accordance with DIN EN ISO 13485 was implemented in order to monitor and continuously improve the structures and processes of the MeDIC.

In cooperation with the University Hospital, the University of Münster operates a Medical Data Integration Center (MeDIC), located at the Institute of Medical Informatics. The MeDIC merges data from health research and care and offers colleagues an optimal research environment with partners of the Medical Informatics Initiative.
The analysis of data from health care leads to new insights and improves medical care. The MeDIC observes local and national data security and data protection regulations. In 2020, a quality management system in accordance with DIN EN ISO 13485 was implemented in order to monitor and continuously improve the structures and processes of the MeDIC.
Partner Representative: Prof. Dr. Julian Varghese
Head of MeDIC: Dr. Michael Storck
Head of MeDIC: Dr. Tobias Brix
Partner within the HiGHmed Consortium:
since 2018
Member of the HiGHmed Association:
since 2020

At the University Hospital of Würzburg, medical care, intensive research and extensive teaching to create cutting-edge medicine. In 19 clinics with polyclinics, three independent polyclinics and four clinical institutes, almost 320,000 patients are treated there each year as outpatients and inpatients. The University Hospital of Würzburg looks back on a history of more than 400 years, making it one of the oldest university hospitals in Germany.
Partner Representative: Prof. Dr. Peter Heuschmann
Head of MeDIC: Maximilian Ertl
Deputy Head of MeDIC: Georg Fette
Partner within the HiGHmed Consortium:
since 2018
Member of the HiGHmed Association:
since 2020

The Carl-Thiem-Hospital has developed into a high-performance medical center and stands for excellent comprehensive medical care. More than 100,000 patients are treated as inpatients and/or outpatients each year. The CTK is an academic teaching hospital of the Charité - Universitätsmedizin Berlin.
Partner Representative: Dr. Steffen Ortmann
Partner within the HiGHmed Consortium:
since 2018
Member of the HiGHmed Association:
since 2020
Own Medical Data Integration Centre (MeDIC) in planning

As a university internationally regarded for its top-level research and innovative teaching concepts, Bielefeld University makes a significant contribution to a progressive and participatory knowledge society. With more than 24,000 students, the University encompasses 14 faculties, covering a broad range of disciplines in the humanities, natural sciences, social sciences and technology.
The Faculty of Medicine OWL has been training its first students since 2021. Its research profile 'People with chronic diseases and disabilities' is future-oriented in the university medical context and highly relevant to society.
The Faculty of Medicine OWL is a pioneer in the development of a competitive research data infrastructure for a university hospital that consists of the university specialist clinics of several hospital owners. The establishment of a joint research data platform enables the development and use of a large variety of medical data and forms an important basis for conducting clinical studies. The expansion into a Medical Data Integration Centre (MeDIC) ensures supraregional connectivity.
Partner Representative: Dr. Laura Dittmar
Member of the HiGHmed Association:
since 2022
Partner within the HiGHmed Consortium:
planned as of 2023
Own Medical Data Integration Centre (MeDIC) in planning

The University of Oldenburg was founded in 1973. This makes it one of the youngest universities in Germany. Its goal is to find answers to the major questions facing society in the 21st century - with top interdisciplinary research and teaching. The university works closely with more than 200 universities worldwide.
The university prepares around 16,000 students for professional life. The spectrum ranges from the humanities and cultural studies to economics, law and social sciences, mathematics, computer science, the natural sciences and medicine.
Partner Representative: Prof. Dr. Antje Wulff
Member of the HiGHmed Association:
since 2022
Partner within the HiGHmed Consortium:
planned as of 2023

The hospital site of Helios University Hospital combines a long history with a long history with innovative, state-of-the-art medicine. The research focus is on "integrative and person-centred healthcare". This includes the comprehensive medical approaches in which not only the disease is in the focus, but always the whole person.
Partner within the HiGHmed Consortium:
As of 2023
Member of the HiGHmed Association:
tba.
Own Medical Data Integration Centre(MeDIC) in planning

The Robert Bosch Hospital is a hospital supported by the Robert Bosch Stiftung at the Bosch Health Campus in Stuttgart. Since 1978, the Robert Bosch Hospital has been one of the academic teaching hospitals of the University of Tübingen.
Member of the HiGHmed Association:
since 2022
Own Medical Data Integration Centre(MeDIC) in planning

The Hannover Medical School (MHH) is one of the most efficient universities in Germany. Whether in research, patient care or teaching, the concept of targeted priority funding has enabled the MHH to secure one of the top places among German university hospitals in recent years.
Partner Representative: Prof. Dr. Michael Marschollek
Head of MeDIC: Dr. Matthias Gietzelt
Deputy Head of MeDIC: Birger Haarbrandt
Partner within the HiGHmed Consortium:
since 2016 (founding location)
Member of the HiGHmed Association:
since 2020
The LIH performs patient-centric translational research with a focus on cancer and immune-related disorders. Its particular interest lies in the immune system as a shared functional module between health and disease.
The LIH team of multidisciplinary researchers embrace collaboration and disruptive technology to advance the understanding of disease mechanisms. Using technology like Artificial Intelligence on real-world patient-derived data, they create disease relevant knowledge, which can be tangibly translated into clinical applications through a bed to bench to bed approach.
As a public biomedical research organisation focused on precision health, the LIH is invested in becoming a leading reference in Europe for the transformation of scientific excellence into meaningful benefits for patients.
Member of the HiGHmed Association:
since 1.11.2022
But we want to do more.
This is why the HiGHmed Association was established as registered association. The HiGHmed Association consolidates the structures and solutions developed as part of the HiGHmed Consortium and aims to establish them permanently and independently beyond the funding period of the project.
In addition, the association supports the governance, coordination, and implementation of other research initiatives and projects within the HiGHmed ecosystem with the objective to quickly transfer new research findings into practice.
This helps to improve prevention, diagnosis, therapy, and care in a sustainable way.

Part of our work is to promote the cooperation with institutions from different areas, including the scientific, political, and industrial sector. The HiGHmed Association hosts seminars, symposia, and workshops with external partners and establishes national and international cooperations and strategic alliances.
The coordination, implementation, and operationalization of central data infrastructure and data management, as well as the definition of and compliance with uniform and transparent standards for the collection, storage and use of data forms the core of our work.
HiGHmed demonstrates how data, information and knowledge can be linked for the benefit of all patients.
HiGHmed

HiGHmed Consortium
The HiGHmed Consortium is working on novel, interoperable solutions in medical informatics with the goal of making medical patient data accessible for clinical research and education.

HiGHmeducation
Under the HiGHmed brand, our academic and private partners combine their expertise in their respective fields to deliver exceptional courses in medical informatics.
HiGHmed Association Projects

NUM RDP
The clinical Routine Data Platform (RDP) establishes a secure, scalable platform offering interoperability in the provision of patient data sourced from clinical routine treatment.
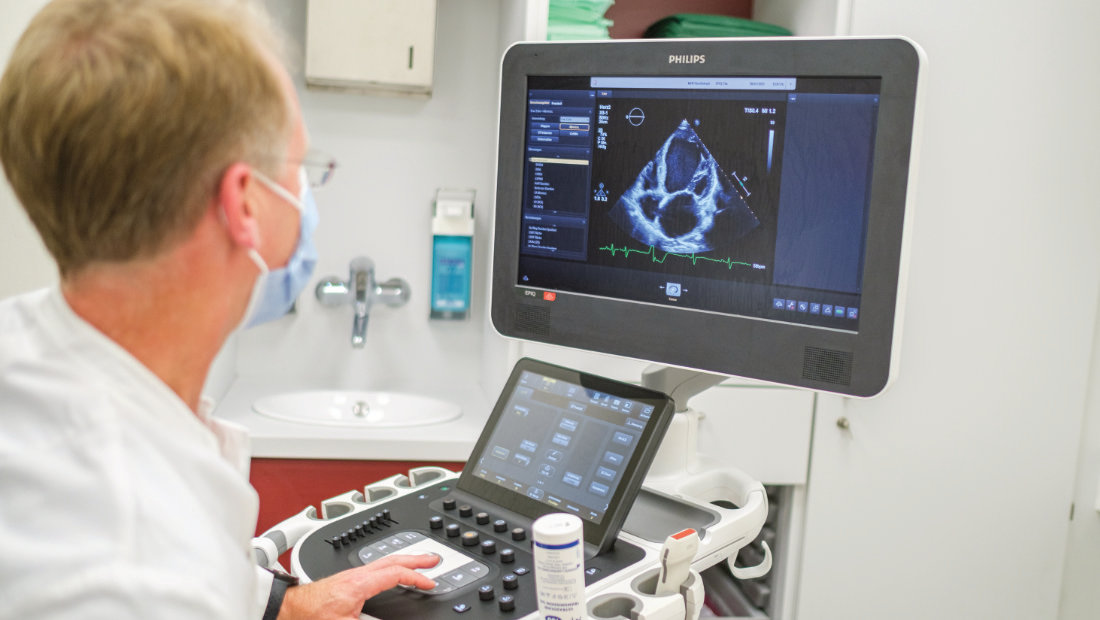
CAEHR
The digital research hub CAEHR - coordinated by the University Medical Center Göttingen - aims to develop and implement a research-compatible electronic patient record (ePA).
CATCOVID
Vaccination and mass testing are the most important tools in tackling the COVID-19 pandemic. In addition, safe and effective drugs are needed to treat those who become sick.

TRANSIT
Vaccination and mass testing are the most important tools in tackling the COVID-19 pandemic. In addition, safe and effective drugs are needed to treat those who become sick.

HEALTH-X dataLOFT
Funded by the German Federal Ministry of Economy and Climate Protection the project aims to implement health data in a legitimized, open and federated dataLOFT platform and to make it accessible in compliance with Gaia-X standards.
HiGHmed Consortium Use Cases (2018-2023)
Use Case Infection Control
Here, a digital system for early detection of potential nosocomial transmission chains is being developed and researched.
Use Case Cardiology
Chronic heart failure affects more than three million people in Germany and is the most common reason for hospitalization.

Use Case Oncology
The UC Oncology focuses on 3 types of cancer that are difficult to treat: tumors of the liver, the bile ducts and the pancreas.
Our Publications
HiGHmed scientists regularly publish their outstanding studies in renowned journals.
Click here for the publication-overview and take a read!



![LIH_Logo[63836]](https://www.highmed.org/hs-fs/hubfs/LIH_Logo%5B63836%5D.png?width=269&height=98&name=LIH_Logo%5B63836%5D.png)


























